 
A central aim of computational structural biology is to fold a protein, i.e., to identify its ( single) Ω m with global minimum F m (out of trillions of microstates) – an unsolved optimization task. MD studies have shown that a molecule will visit a localized well only for a very short time (as short as several fs) while staying for a much longer time within a microstate, meaning that the microstates are of a greater physical significance than the localized wells. 1) a microstate can be represented by a sample (trajectory) generated by a local MD simulation (e.g., the α-helical region of a peptide, see further discussion in II.4 below). More specifically, this surface is “decorated” by a tremendous number of localized energy wells and “wider” ones that are defined over microstates (regions Ω m), each consisting of many localized wells ( Fig. While the difficulty in calculating the absolute S ( F) discussed above is common to all systems, biological macromolecules such as peptides and proteins, are particularly challenging due to their rugged potential energy surface, E( x). II.3 Microstates and intermediate flexibility The present article constitutes a substantial extension of a concise review appeared recently. We summarize here mainly recent developments in this area of research where the emphasis is on methodology issues and less on applications. While significant progress has been made (see reviews in ), in many cases the efficiency (or accuracy) of existing methods is unsatisfactory and the need for new ideas has kept this field highly active. However, calculation of F( S) by computer simulation is extremely difficult, and considerable attention has thus been devoted in the last 50 years to this subject. The free energy defines the binding affinities of protein-protein and protein-ligand interactions, it also quantifies many other important processes such as enzymatic reactions, electron transfer, ion transport through membranes, and the solvation of small molecules.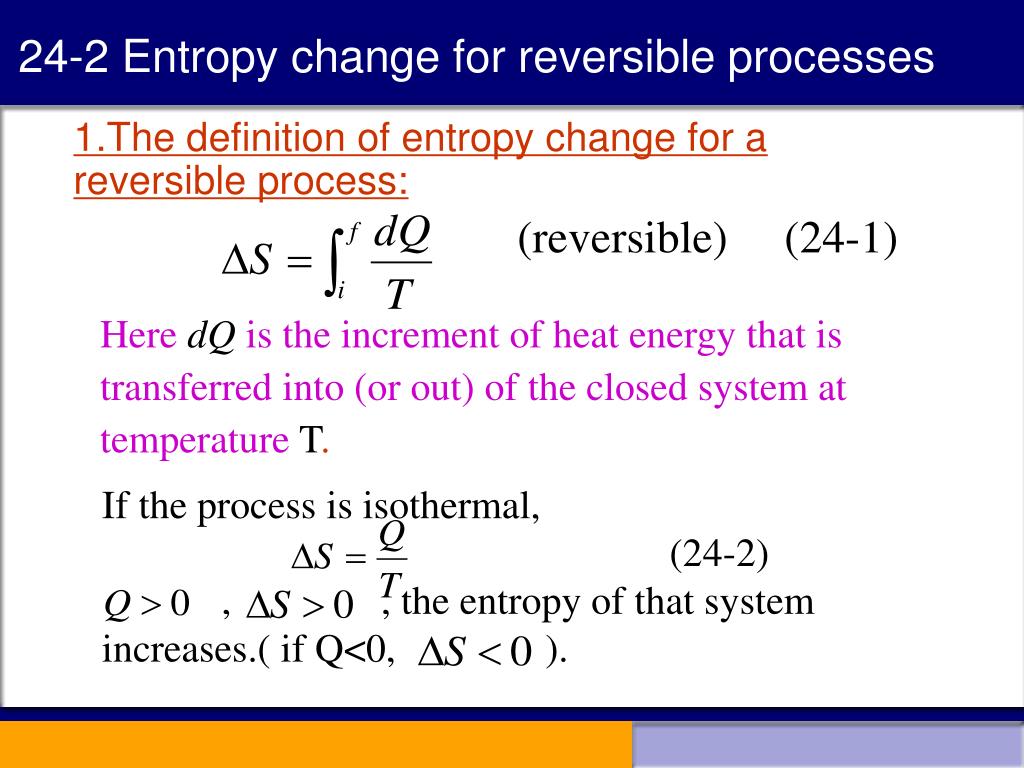 
F constitutes the criterion of stability, which is essential for studying the structure and function of peptides, proteins, nucleic acids, and other biological macromolecules. S is a measure of order where changes in the S of water lead to the hydrophobic interaction – the main driving force in protein folding. The fact that the absolute value of specific entropy is unknown is not a problem because it is the change in specific entropy (Δs) and not the absolute value that is important in practical problems.The absolute entropy, S and the absolute Helmholtz free energy, F (or G – Gibbs free energy) are fundamental quantities in statistical mechanics with a special importance in structural biology. 
For example, the specific entropy of water or steam is given using the reference that the specific entropy of water is zero at 32☏. Also, like enthalpy, the entropy of a substance is given with respect to some reference value. Like enthalpy, entropy cannot be measured directly. Entropy is represented by the letter S and can be defined as ΔS in the following relationships. 
Entropy is sometimes referred to as a measure of the inability to do work for a given heat transferred. Because entropy tells so much about the usefulness of an amount of heat transferred in performing work, the steam tables include values of specific entropy (s = S/m) as part of the information tabulated. Entropy quantifies the energy of a substance that is no longer available to perform useful work. Because entropy is a property, changes in it can be determined by knowing the initial and final conditions of a substance. > Thermodynamics Directory | Heat Transfer DirectoryĮntropy Definition - Thermodynamic PropertiesĮntropy (S) is a property of a substance, as are pressure, temperature, volume, and enthalpy. Entropy Definition and Equation Thermodynamics 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
